A “primer” on cannabis sequencing and testing
Advances in gene technology have led to a revolution in cannabis breeding and cultivation. CRISPR, a relatively recent and yet widely popular gene-editing method, is quickly being integrated into the tool belt of technicians across industries. [1] While colloquially the technique is described as “cut and paste” editing, it’s not quite that simple, and, in fact, its discovery was predicated by decades of research into DNA, the building blocks of our genes. So let’s take a look at how genetic structure was first resolved.
As many know, Rosalind Franklin, James Watson, and Francis Crick first described their double-helix model of DNA in 1953. [2] Their work kick-started the field of human molecular genetics, which culminated in mapping the human genome in 2001. [3]But before the human genome could be mapped, scientists first needed to figure out how to “decode” genetic information and read the sequence of base pairs (adenine, guanine, cytosine, and thymine) that make up a strand of DNA.
To properly “read” a sample of DNA, the genetic material must be amplified to produce many identical copies. Early DNA sequencing methods involved insertion of a gene of interest into living bacterial cells; however, this process was very time-consuming. In the 1980s, a new technique called polymerase chain reaction, or PCR, was developed that enabled the production of millions of copies of a DNA sequence within just a few hours. [4] While alternate methods of genetic sequencing have been developed since then, PCR remains a popular technique still used in genetic labs worldwide.
It’s worth noting that PCR was developed in part due to the mind-expanding effects of LSD. While Kay Mullis, PhD, a scientist at a biotech company, was working on automated DNA replication, he happened to meet Albert Hoffman, PhD, who had invented LSD. Through substance-induced exploration and relentless testing, PCR was born.
PCR is implemented across a series of steps, the first of which involves denaturation, or separation of the complementary DNA strands using high temperatures. [5] The material is then cooled and primers, or short sequences of base pairs complementary to pieces of the sequence to be replicated, bind to specific segments of each strand (“annealing”). During the third step, DNA polymerase, the enzyme that elongates a DNA chain, creates a complementary copy of the target sequence. This cycle is repeated 40 or 50 times to create enough copies of DNA for evaluation.
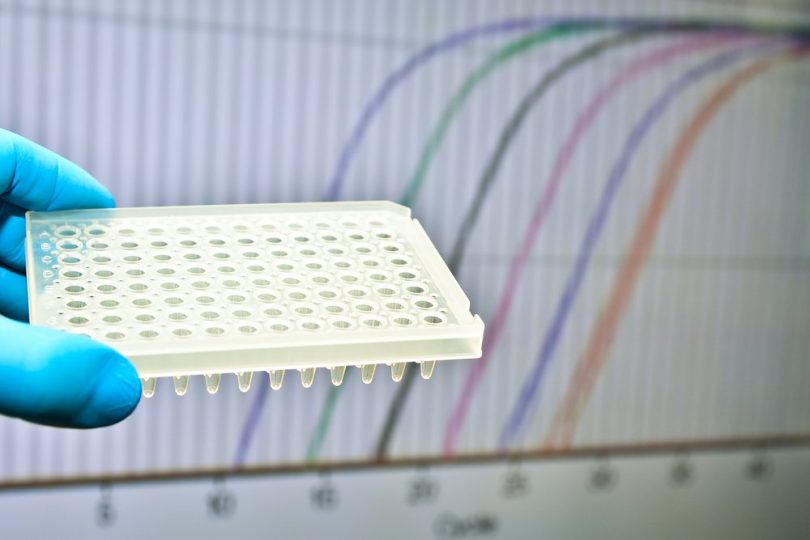
Quantitative PCR, or qPCR, is a type of PCR that uses florescence dyes to label DNA in “real-time,” so that DNA sequences can be copied and quantified across each cycle. [5] The data from this method can be visualized by plotting fluorescence level against cycle number compared to a standard value.
Gene sequencing technology is used in cannabis breeding and testing. First, qPCR can identify a particular cultivar or hybrid. [6] Based on this information, breeders can select for the most desirable traits of the plant, including type and level of cannabinoid, terpenes, color, aroma, and behavioral effects, to produce a highly tailored product. What once took years of crossing cultivars is now much faster and more efficient with genetic technology.
qPCR also plays a very important role in cannabis testing. Before cannabis products can be safely consumed, they should be tested for potentially harmful microorganisms, such as bacteria and fungi. [7] Genetic sequencing via qPCR can both detect and identify pathogens within a few hours with high sensitivity and specificity. This type of testing is essential to protect cannabis users from ingesting unsafe products and is of the utmost importance for patients who use medical cannabis and are immunocompromised. [8]
As the industry continues to thrive, cannabis breeding and testing must remain an important facet that drives the creation of consistent, safe, high-quality products by utilizing the appropriate genetic technology.
References
- Ophinni, Y., Inoue, M., Kotaki, T., Kameoka, M., “CRISPR/Cas9 System Targeting Regulatory Genes of HIV-1 Inhibits Viral Replication in Infected T-cell Cultures”, Scientific Reports, 2018, Volume 8, pg. 1-12. (impact factor: 4.122; cited by: 4)
- Watson, J.D., Crick, H.C., “Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid”, Nature, Volume 171, pg. 737-738. (impact factor: 41.577; cited by: 15,410)
- Lander, E.S., Linton, L.M., Birren, B, et al., “Initial Sequencing and Analysis of the Human Genome”, Nature, 2001, Volume 409, pg. 860-921.(impact factor: 41.577; cited by: 23,060)
- Mullis, K., Faloona, F., Scharf, S., Saiki, R., Horn, G., Erlich, H., “Specific Enzymatic Amplification of DNA In Vitro: The Polymerase Chain Reaction”, Cold Spring HarbSymp Quant Biol, 1986, Volume 51, pg. 263-273. (impact factor: N/A; cited by: 4,858)
- Garibyan, L.,Avashia, N., “Research Techniques Made Simple: Polymerase Chain Reaction (PCR)”, J Invest Dermatol, 2013, Volume 133, pg. 1-8. (impact factor: 6.448; cited by: 163)
- Mendoza, M.A., Mills, D.K.,Lata, H., Chandra, S.,ElSohly, M.A., Almirall, J.R., “Genetic Individualization of Cannabis Sativa by a Short Tandem Repeat Multiplex System”, Anal BioanalChem, 2009, Volume 393, pg. 719-726. (impact factor: 3.578; cited by: 24)
- McKernan, K., Spangler, J., Zhang, L., et al., “Cannabis Microbiome Sequencing Reveals Several Mycotoxic Fungi Native to Dispensary Grade Cannabis Flowers”, F1000Res, 2016, Volume 44, pg. 1-25. (impact factor: N/A; cited by: 10)
- Thompson, G.R., Tuscano, J.M., Dennis, M., et al., “A Microbiome Assessment of
Medical Marijuana”, ClinMicrobiol Infect, 2017, Volume 23, pg. 269-270. (impact factor: 5.349; cited by: 14)